 -
Volume 85,
Issue 7,
2004
-
Volume 85,
Issue 7,
2004
Volume 85, Issue 7, 2004
- Animal
-
- RNA viruses
-
-
IL8 release, tight junction and cytoskeleton dynamic reorganization conducive to permeability increase are induced by dengue virus infection of microvascular endothelial monolayers
More LessPermeability alterations of microvascular endothelia may be a factor in the plasma leakage produced by dengue virus infection. Confluent monolayers of the human dermal microvascular endothelial cell line HMEC-1 were utilized as an experimental model to study the cellular responses induced by the virus. Infected monolayers showed increased permeability for [3H]mannitol, but no changes were observed for 4–70 kDa dextrans at 48 h post-infection (p.i.), a time at which viral titres reached maximal values and 40 % of the cells expressed viral proteins. A further increase in permeability occurred at 72 h, still without evident cytopathic effects on the monolayer. Coinciding with this, actin was reorganized in the infected cells and the tight junction protein occludin was displaced to the cytoplasm. Increments in the thickness of stress fibres and focal adhesions were observed in uninfected cells neighbouring infected cells. Culture medium from infected monolayers induced permeability changes and thickening of actin-containing structures in control cultures that resembled those observed 48 h p.i. Interleukin (IL) 8 was found in culture medium at concentrations ranging from 20 to 100 pg ml−1. Neutralizing antibodies against IL8 partially inhibited the changes produced by the culture medium as well as those induced by addition of IL8. Genistein inhibited the effect of the culture medium and the phosphorylation of proteins associated with focal adhesions and indicated the participation of tyrosine kinases. These findings suggest that IL8 production by infected monolayers contributes to the virus-induced effect on the cytoskeleton and tight junctions and thereby modifies transendothelial permeability.
-
-
-
A novel protein expression strategy using recombinant bovine respiratory syncytial virus (BRSV): modifications of the peptide sequence between the two furin cleavage sites of the BRSV fusion protein yield secreted proteins, but affect processing and function of the BRSV fusion protein
More LessThe bovine respiratory syncytial virus (BRSV) fusion (F) protein is cleaved at two furin cleavage sites, which results in generation of the disulfide-linked F1 and F2 subunits and release of an intervening peptide of 27 aa (pep27). A series of mutated open reading frames encoding F proteins that lacked the entire pep27, that contained an arbitrarily chosen 23 aa sequence instead of pep27 or in which pep27 was replaced by the amino acid sequences for the bovine cytokines interleukin 2 (boIL2), interleukin 4 (boIL4) or gamma interferon (boIFN-γ) was constructed. Transient expression experiments revealed that the sequence of the intervening peptide influenced intracellular transport, maturation of the F protein and F-mediated syncytium formation. Expression of boIL2, boIL4 or boIFN-γ in place of pep27 resulted in secretion of the cytokines into the culture medium. All mutated F proteins except the boIFN-γ-containing variant could be expressed by and were functional for recombinant BRSV. Characterization of the cell culture properties of the recombinants demonstrated that the amino acid sequence between the two furin cleavage sites affected entry into target cells, direct spreading of virions from cell to cell and virus growth. Secretion of boIL2 and boIL4 into the medium of cells infected with the respective recombinants demonstrated that the F protein can be used to express secreted heterologous bioactive peptides or (glyco)proteins, which might be of interest for the development of novel RSV vaccines.
-
-
-
Protective lactogenic immunity conferred by an edible peptide vaccine to bovine rotavirus produced in transgenic plants
More LessVaccines produced in transgenic plants constitute a promising alternative to conventional immunogens, presenting the possibility of stimulating secretory and systemic immunity against enteric pathogens when administered orally. Protection against enteric pathogens affecting newborn animals requires, in most cases, the stimulation of lactogenic immunity. Here, the group presents the development of an experimental immunogen based on expression of an immunorelevant peptide, eBRV4, of the VP4 protein of bovine rotavirus (BRV), which has been described as harbouring at least one neutralizing epitope as well as being responsible for the adsorption of the virus to epithelial cells. The eBRV4 epitope was efficiently expressed in transgenic alfalfa as a translational fusion protein with the highly stable reporter enzyme β-glucuronidase (βGUS), which served as a carrier, stabilized the synthesized peptide and facilitated screening for the higher expression levels in plants. Correlation of expression of the eBRV4 epitope in plants with those presenting the highest βGUS activities was confirmed by a Western blot assay specific for the BRV peptide. The eBRV4 epitope expressed in plants was effective in inducing an anti-rotavirus antibody response in adult female mice when administered either intraperitoneally or orally and, more importantly, suckling mice born from immunized female mice were protected against oral challenge with virulent rotavirus. These results demonstrate the feasibility of inducing lactogenic immunity against an enteric pathogen using an edible vaccine produced in transgenic plants.
-
-
-
Antibodies specific for hypervariable regions 3 to 5 of the feline immunodeficiency virus envelope glycoprotein are not solely responsible for vaccine-induced acceleration of challenge infection in cats
More LessIn a previous vaccination study in cats, the authors reported on accelerated feline immunodeficiency virus (FIV) replication upon challenge in animals vaccinated with a candidate envelope subunit vaccine. Plasma transfer studies as well as antibody profiles in vaccinated cats indicated a causative role for antibodies directed against the hypervariable regions HV3, HV4 and HV5 (HV3–5) of the envelope glycoprotein. The present study was designed to investigate further the contribution of antibodies in envelope vaccine-induced acceleration of FIV infection. To this end, regions HV3–5 of the envelope glycoprotein were deleted from the original vaccine, thus addressing the contributing role of antibodies directed against these hypervariable regions. Interestingly, this approach did not prevent acceleration of challenge infection. Analysis of the antibody responses in the respective groups suggested that removal of HV3–5 redirected the humoral immune response towards other regions of the envelope glycoprotein, indicating that these regions can also induce antibodies that accelerate virus replication.
-
-
-
Feline immunodeficiency virus subtypes A, B and C and intersubtype recombinants in Ontario, Canada
More LessF. Reggeti and D. BienzleKnowledge of the geographical distribution of feline immunodeficiency virus (FIV) subtypes is important for understanding different disease courses and for vaccine design. Intersubtype recombination may develop in areas where more than one subtype is prevalent and has the potential to create new transmittable variants with novel pathogenic properties. In this study, 40 FIV-positive DNA samples were classified by sequence analysis of the LTR–gag region. Phylogenetic analysis indicated that 32 Canadian FIV isolates clustered with previously identified subtypes A, B and C and that subtype A was most frequent in Ontario. Four strains with inconsistent clade assignment were further analysed by sequencing of the env–LTR regions. Comparisons of phylogenetic trees constructed from the two different regions of the genome and analysis of similarities to reference sequences yielded classification of three samples as A/B and one as A/C intersubtype recombinants. Although the A/B recombinant samples were obtained from unrelated cats in geographically disparate regions, a common breakpoint was consistently identified within gag. In addition, there was no evidence of co-infection with parental strains of subtypes A and B as indicated by PCR-based limiting dilution assays, although these assays allowed for the identification of two different recombinant viruses co-existing in one sample. Both sequences contained the same breakpoint. These findings suggested that a new circulating recombinant FIV may be enzootic in Ontario.
-
-
-
Full-length open reading frame of a recombinant hepatitis C virus strain from St Petersburg: proposed mechanism for its formation
More LessThe full-length ORFs for the hepatitis C virus recombinant RF1_2k/1b (N687) and the non-recombinant 1b strain N589 were sequenced. A single recombination point was found and the sizes of the genes (C, E1, E2, p7, NS2, NS3, NS4 and NS5) were according to the parental subtypes. The PKR-eIF2α phosphorylation site homology domain sequence of the E2 protein was identical to those of genotype 2 strains, while the IFN-α-sensitivity-determining region of the NS5A protein was identical to those of interferon-resistant 1b strains. For the parental strains, two hairpin structures, HS1 and HS2, were predicted for the plus-strand up- and downstream of the crossover site, which were not present in the recombinant strain. HS2 shared similarity with the motif1 hairpin of turnip crinkle virus RNA that binds to the RNA-dependent RNA polymerase and facilitates 3′-terminal extension during recombination. This study suggests that RF1_2k/1b has emerged by homologous recombination during minus-strand synthesis via template switching because of constraints imposed by the HS1 hairpin of the 3′-parental genome.
-
-
-
Evolution of naturally occurring 5′ non-translated region variants of hepatitis C virus genotype 1b in selectable replicons
More LessQuasispecies shifts are essential for the development of persistent hepatitis C virus (HCV) infection. Naturally occurring sequence variations in the 5′ non-translated region (NTR) of the virus could lead to changes in protein expression levels, reflecting selective forces on the virus. The extreme 5′ end of the virus' genome, containing signals essential for replication, is followed by an internal ribosomal entry site (IRES) essential for protein translation as well as replication. The 5′ NTR is highly conserved and has a complex RNA secondary structure consisting of several stem–loops. This report analyses the quasispecies distribution of the 5′ NTR of an HCV genotype 1b clinical isolate and found a number of sequences differing from the consensus sequence. The consensus sequence, as well as a major variant located in stem–loop IIIa of the IRES, was investigated using self-replicating HCV RNA molecules in human hepatoma cells. The stem–loop IIIa mutation, which is predicted to disrupt the stem structure, showed slightly lower translation efficiency but was severely impaired in the colony formation of selectable HCV replicons. Interestingly, during selection of colonies supporting autonomous replication, mutations emerged that restored the base pairing in the stem–loop. Recloning of these altered IRESs confirmed that these second site revertants were more efficient in colony formation. In conclusion, naturally occurring variants in the HCV 5′ NTR can lead to changes in their replication ability. Furthermore, IRES quasispecies evolution was observed in vitro under the selective pressure of the replicon system.
-
-
-
Dominant negative effect of wild-type NS5A on NS5A-adapted subgenomic hepatitis C virus RNA replicon
More LessAn efficient model is currently used to study hepatitis C virus (HCV) replication in cell culture. It involves transfection in Huh7, a hepatoma-derived cell line, of an antibiotic (neomycin) selectable HCV subgenomic replicon encoding the non-structural (NS) proteins from NS3 to NS5B. However, strong and sustained replication is achieved only on the appearance of adaptive mutations in viral proteins. The most effective of these adaptive mutations are concentrated mainly in NS5A, not only into the original Con1 but also in the recently established HCV-BK and HCV-H77 isolate-derived replicons. This suggests that the expression of wild-type (wt) NS5A may not allow efficient HCV RNA replication in cell culture. With the use of a β-lactamase reporter gene as a marker for HCV replication and TaqMan RNA analysis, the replication of different HCV replicons in cotransfection experiments was investigated. Comparing wt with NS5A-adapted replicons, the strong evidence accumulated showed that the expression of wt NS5A was actually able to inhibit the replication of NS5A-adapted replicons. This feature was characterized as a dominant negative effect. Interestingly, an NS5B (R2884G)-adapted replicon, containing a wt NS5A, was dominant negative on an NS5A-adapted replicon but was not inhibited by the original Con1 replicon. In conclusion, these studies revealed that the original wt Con1 replicon is not only incompetent for replication in cell culture, but is also able to interfere with NS5A-adapted replicons.
-
-
-
Inhibition of influenza virus matrix (M1) protein expression and virus replication by U6 promoter-driven and lentivirus-mediated delivery of siRNA
More LessSmall interfering RNA (siRNA)-induced RNA degradation has been used recently as an antivirus agent to inhibit specific virus replication. This report shows that 21 nt duplexes of siRNA of the influenza virus M gene can cause specific inhibition of influenza virus matrix (M1) protein expression in transfected 293T cells. Furthermore, it is shown that a lentivirus vector can be used to effectively deliver M gene siRNAs into Madin–Darby canine kidney cells and can cause specific inhibition of M1 protein expression and influenza virus replication. Therefore, lentivirus-mediated delivery of siRNA and gene silencing can be used in studying the specific functions of virus genes in replication and may have a potential therapeutic application.
-
-
-
Identification of an amino acid residue on influenza C virus M1 protein responsible for formation of the cord-like structures of the virus
More LessInfluenza C virus-like particles (VLPs) have been generated from cloned cDNAs. A cDNA of the green fluorescent protein (GFP) gene in antisense orientation was flanked by the 5′ and 3′ non-coding regions of RNA segment 5 of the influenza C virus. The cDNA cassette was inserted between an RNA polymerase I promoter and terminator of the Pol I vector. This plasmid DNA was transfected into 293T cells together with plasmids encoding virus proteins of C/Ann Arbor/1/50 or C/Yamagata/1/88. Transfer of the supernatants of the transfected 293T cells to HMV-II cells resulted in GFP expression in the HMV-II cells. The quantification of the GFP-positive HMV-II cells indicated the presence of approximately 106 VLPs (ml supernatant)−1. Cords 50−300 μm in length were observed on transfected 293T cells, although the cords were not observed when the plasmid for M1 protein of C/Ann Arbor/1/50 was replaced with that of C/Taylor/1233/47. A series of transfection experiments with plasmids encoding M1 mutants of C/Ann Arbor/1/50 or C/Taylor/1233/47 showed that an amino acid at residue 24 of the M1 protein is responsible for cord formation. This finding provides direct evidence for a previous hypothesis that M1 protein is involved in the formation of cord-like structures protruding from the C/Yamagata/1/88-infected cells. Evidence was obtained by electron microscopy that transfected cells bearing cords produced filamentous VLPs, suggesting the potential role of the M1 protein in determining the filamentous/spherical morphology of influenza C virus.
-
-
-
The X protein of Borna disease virus regulates viral polymerase activity through interaction with the P protein
More LessBorna disease virus polymerase activity is negatively regulated by the viral X protein. Using a virus minireplicon system it was found that all X mutants that no longer interacted with the viral P protein failed to exhibit significant inhibitory activity. The action of X could further be neutralized by expression of a P fragment that contained the X interaction domain but lacked all domains known to mediate interaction with other viral proteins. X thus appears to regulate the activity of the Borna disease virus polymerase by targeting the polymerase cofactor P.
-
-
-
Comparison of the effects of RNase-negative and wild-type classical swine fever virus on peripheral blood cells of infected pigs
More LessElimination of the RNase activity of classical swine fever virus (CSFV) glycoprotein Erns was previously shown to result in virus attenuation. Specific reduction of B cell numbers in the peripheral blood, a typical symptom of CSFV infection in pigs, was not detected on infection with the RNase-negative mutant C-H346Δ [ Meyers et al. (1999). J Virol 73, 10224–10235 ]. The present report shows that this feature is restricted to this specific virus mutant, and does not represent a general property of RNase-negative CSFV. The effects induced by infection with two other RNase-negative and wild-type (wt) CSFV strains on the composition of peripheral blood cells have been further analysed. For all viruses, not only general leukopenia but also a reduction of different subsets of leukocytes (T-lymphocytes, monocytes and granulocytes) was detected. Similar to the results with B-cells, no significant differences with regard to changes in cell number were determined for RNase-negative mutants and wt virus during the initial phase of infection. Later, the values returned to pre-infection levels for the mutants, but stayed at low levels in the wt virus-infected animals. A major difference was reflected in the virus load of the infected animals, which was dramatically higher for pigs infected with wt CSFV, so that reduction of the virus load represents a further marker for attenuation resulting from RNase destruction. Attenuation was also detectable for the RNase-negative mutant C-W300G, which showed rapid reversion to the wt sequence within the infected pig. The prevention of fatal disease after infection with C-W300G is apparently determined during the short time between infection and reversion, as the virus revertant reisolated from infected pigs was shown to be virulent when used for infection in a follow-up study. Reversion of C-W300G was also detected in tissue culture during passage on swine testis epithelioid cells and porcine transformed kidney (MAX) cells, whereas the mutation was stable when SK6 or 38A1D cells were tested.
-
-
-
Identification and analysis for cross-reactivity among hantaviruses of H-2b-restricted cytotoxic T-lymphocyte epitopes in Sin Nombre virus nucleocapsid protein
More LessSin Nombre virus (SNV) causes hantavirus pulmonary syndrome (HPS), with a high rate of mortality in humans who are infected by the transmission of virus from the natural rodent host. In humans, cytotoxic T lymphocytes (CTL) specific for SNV appear to play an important role in the pathogenicity of HPS. There is a correlation between the frequencies of SNV-specific CTLs and the severity of HPS disease. In order to create a mouse model to study the role of SNV-specific T cells in vivo, T cell responses to SNV nucleocapsid (N) protein in B6.PL Thy1 a/Cy mice (H-2b) immunized with plasmid DNA or recombinant vaccinia virus expressing SNV N protein were examined. Four peptides, NC94–101, NC175–189, NC217–231 and NC331–345, were recognized by CD8+ T cells in CTL and ELISPOT assays in SNV N-immunized mice. Interestingly, two of these epitopes are located in the central region of the SNV N protein, where several human CD8+ T-cell epitopes have been defined in Puumala virus and SNV. CTL lines specific for these four epitopes were cross-reactive to corresponding Puumala virus peptides, but only one of them was cross-reactive to Hantaan virus peptides. These results will enable the analysis of the roles of CTL in immunopathology of HPS in experimental mouse models of HPS.
-
-
-
Tax protein of human T-cell leukaemia virus type 1 induces interleukin 17 gene expression in T cells
More LessTax protein of human T-cell leukaemia virus type 1 (HTLV-1) induces the expression of several cellular genes that are involved in T cell activation and proliferation. In this study, it was observed that Tax upregulated the expression of human interleukin 17 (IL17), a cytokine mainly produced by activated CD4+ memory T cells. Indeed, IL17 mRNA was highly expressed in HTLV-1-infected T cells as well as in Tax-expressing Jurkat T cells, whereas it was not detectable in HTLV-1-negative T cell lines. The clinical relevance of these observations was further demonstrated by quantitative assessment of IL17 expression in lymphocytes isolated from one HTLV-1-infected patient. To define the transcriptional activation of the IL17 gene by Tax, the 5′-flanking region of this gene was cloned and a reporter gene analysis performed. The presence of a Tax-responsive region spanning 614 bp upstream of the initiation start site was identified, in HeLa as well as in Jurkat cells, stimulated with phorbol myristate acetate and Ca2+ ionophore. Finally, Tax mutants were used to show that the transcriptional activation of the IL17 promoter by Tax was dependent on the CREB/ATF pathway. As IL17 upregulates the expression of several pro-inflammatory cytokines, these observations provide new insights into the involvement of the Tax protein in the pathophysiology of HTLV-1-associated inflammatory disorders.
-
-
-
Characterization of in vitro and in vivo phenotypes of poliovirus type 1 mutants with reduced viral protein synthesis activity
More LessSabin vaccine strains of poliovirus (PV) contain major attenuation determinants in the internal ribosomal entry site (IRES), an area that directs viral protein synthesis. To examine the effect of reduced viral protein synthesis on PV neurovirulence, spacer sequences, consisting of short open reading frames of different lengths, were introduced between the IRES and the initiation codon of viral polyprotein, resulting in PV mutants with reduced viral protein synthesis. These PV mutants had a viral protein synthesis activity 8·8–55 % of that of the parental Mahoney strain as measured in HeLa S3 cells. Only viruses with more than 28 % of the wild-type activity had intact spacer sequences following plaque purification. Mutants with 17 % or 21 % of the wild-type activity were unstable and a mutant with 8·8 % was lethal. The neurovirulence of PV mutants was evaluated in transgenic mice carrying the human PV receptor gene. In this test, mutants with more than 28 % of the wild-type activity remained neurovirulent, while a mutant with 17 % of wild-type activity exhibited a partially attenuated phenotype. This mutant stably replicated in the spinal cord; however, the stability was severely affected during the course of virus infection from the cerebrum to the spinal cord. These results suggest that reduced viral protein synthesis activity as measured in cultured cells (17–55 % of the wild-type activity) is not the main determinant of PV attenuation.
-
-
-
Human immunodeficiency virus 1 downregulates cell surface expression of the non-classical major histocompatibility class I molecule HLA-G1
More LessHuman immunodeficiency virus 1 (HIV-1) downregulates cell surface expression of HLA-A and HLA-B but not HLA-C or HLA-E to ultimately escape immune defences. Here, it is shown that cell surface expression of the non-classical HLA-G1 is also downregulated by HIV-1, by using co-transfection experiments and infection with cell-free HIV-1 of HLA-G1-expressing U87 glioma cells or macrophages in primary culture. Moreover, co-transfection experiments using proviruses deleted in either nef or vpu or plasmids encoding HIV-1 Nef and Vpu mixed together with a HLA-G1-expressing construct demonstrated that HLA-G1 downregulation is Nef-independent and Vpu-dependent, contrasting with the Nef- and Vpu-dependent HLA-A2 downregulation. Together, these results show that the decrease of HLA-A2 and HLA-G1 caused by HIV-1 occurs through distinct mechanisms.
-
-
-
Loss of virus-specific CD4+ T cells with increases in viral loads in the chronic phase after vaccine-based partial control of primary simian immunodeficiency virus replication in macaques
More LessVirus-specific cellular immune responses play an important role in the control of immunodeficiency virus replication. However, preclinical trials of vaccines that induce virus-specific cellular immune responses have failed to contain simian immunodeficiency virus (SIV) replication in macaques. A defective provirus DNA vaccine system that efficiently induces virus-specific CD8+ T-cell responses has previously been developed. The vaccinated macaques showed reduced viral loads, but failed to contain SIVmac239 replication. In this study, macaques that showed partial control of SIV replication were followed up to see if or how they lost this control in the chronic phase. Two of them showed increased viral loads about 4 or 8 months after challenge and finally developed AIDS. Analysis of SIV-specific T-cell levels by detection of SIV-specific gamma interferon (IFN-γ) production revealed that these two macaques maintained SIV-specific CD8+ T cells, even after loss of control, but lost SIV-specific CD4+ T cells when plasma viral loads increased. The remaining macaque kept viral loads at low levels and maintained SIV-specific CD4+ T cells, as well as CD8+ T cells, for more than 3 years. Additional analysis using macaques vaccinated with a Gag-expressing Sendai virus vector also found loss of viraemia control, with loss of SIV-specific CD4+ T cells in the chronic phase of SIV infection. Thus, SIV-specific CD4+ T cells that were able to produce IFN-γ in response to SIV antigens were preserved by the vaccine-based partial control of primary SIV replication, but were lost with abrogation of control in the chronic phase.
-
-
-
Strand transfer to the 5′ part of a tRNA as a mechanism for retrovirus patch-repair recombination in vivo
More LessBy screening for marker-cassette deletion mutants of a murine leukaemia virus-based replication-competent vector, two occurrences of tRNA sequence patch insertions were identified. In one of the cases, 28 nucleotides from the 5′ end of tRNALys4 were inserted in the plus-strand orientation, which points to a novel strand-transfer mechanism to tRNAs during reverse transcriptase-mediated retroviral recombination.
-
-
-
Sequences of flavivirus-related RNA viruses persist in DNA form integrated in the genome of Aedes spp. mosquitoes
More LessFlavivirus-related sequences have been discovered in the dsDNA genome of Aedes albopictus and Aedes aegypti mosquitoes, demonstrating for the first time an integration into a eukaryotic genome of a multigenic sequence from an RNA virus that replicates without a recognized DNA intermediate. In the Aedes albopictus C6/36 cell line, an open reading frame (ORF) of 1557 aa with protease/helicase and polyprotein processing domains characteristic of flaviviruses was identified. It is closely related to NS1–NS4A genes of the Cell Fusing Agent and Kamiti River virus and the corresponding mRNAs were detected. Integrated sequences homologous to the envelope, NS4B and polymerase genes of flaviviruses were identified. Overall, approximately two-thirds of a flavivirus-like genome were characterized. In the Aedes aegypti A20 cell line, a 492 aa ORF related to the polymerase of the Cell Fusing Agent and Kamiti River virus was identified. These flavivirus-related integrated DNA sequences were detected in laboratory-bred and wild Aedes albopictus and Aedes aegypti mosquitoes, demonstrating that their discovery is not an artefact resulting from the manipulation of mosquito cell lines, since they exist under natural conditions. This finding has major implications regarding evolution, as it represents an entirely different mechanism by which genetic diversity may be generated in eukaryotic cells distinct from accepted processes.
-
-
-
Sialidase, receptor-binding and fusion-promotion activities of Newcastle disease virus haemagglutinin–neuraminidase glycoprotein: a mutational and kinetic study
More LessMutations were generated in residues at the putative catalytic site of the haemagglutinin–neuraminidase (HN) protein of Newcastle disease virus Clone 30 strain (Arg498, Glu258, Tyr262, Tyr317 and Ser418) and their effects on its three associated activities were studied. Expression of the mutant proteins at the surface of HeLa cells was similar to that of the wild-type. Sialidase, receptor-binding and fusion-promotion activities were affected to different degrees for all mutants studied. Mutant Arg498Lys lost most of its sialidase activity, although it retained most of the receptor-binding activity, suggesting that, for the former activity, besides the presence of a basic residue, the proximity to the substrate molecule is also important, as Lys is shorter than Arg. Proximity also seems to be important in substrate recognition, since Tyr262Phe retained most of its sialidase activity while Tyr262Ser lost most of it. Also, Ser418Ala displayed most of the wild-type sialidase activity. However, a kinetic and thermodynamic study of the sialidase activity of the Tyr262Ser and Ser418Ala mutants was performed and revealed that the hydroxyl group of these residues also plays an important role in catalysis, since such activity was much less effective than that of the wild-type and these mutations modified their activation energy for sialidase catalysis. The discrepancy of the modifications in sialidase and receptor-binding activities in the mutants analysed does not account for the topological coincidence of the two sites. These results also suggest that the globular head of HN protein may play a role in fusion-promotion activity.
-
- DNA viruses
-
-
Internal cleavage of the human cytomegalovirus UL37 immediate-early glycoprotein and divergent trafficking of its proteolytic fragments
More LessThe human cytomegalovirus UL37 gene encodes at least three isoforms, which share N-terminal UL37 exon 1 (UL37x1) sequences. UL37 proteins traffic dually into the endoplasmic reticulum (ER) and to mitochondria. Trafficking of the UL37 glycoprotein (gpUL37) in relation to its post-translational processing was investigated. gpUL37 is internally cleaved in the ER and its products traffic differentially. Its C-terminal fragment (UL37COOH) is ER-localized and N-glycosylated. Unlike conventional ER signal sequences, its N-terminal (
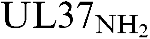 ) fragment is stable and traffics to mitochondria. Inhibition of N-glycosylation did not block pUL37 cleavage and dramatically decreased the levels of
) fragment is stable and traffics to mitochondria. Inhibition of N-glycosylation did not block pUL37 cleavage and dramatically decreased the levels of  but not of UL37COOH. pUL37M, which differs from gpUL37 by the lack of residues 178–262 and hence the UL37x3 consensus signal peptidase cleavage site, traffics into the ER and mitochondria, but is neither cleaved nor N-glycosylated. This finding of a relationship between ER processing and mitochondrial importation of UL37 proteins is unique for herpesvirus proteins.
but not of UL37COOH. pUL37M, which differs from gpUL37 by the lack of residues 178–262 and hence the UL37x3 consensus signal peptidase cleavage site, traffics into the ER and mitochondria, but is neither cleaved nor N-glycosylated. This finding of a relationship between ER processing and mitochondrial importation of UL37 proteins is unique for herpesvirus proteins.
-
-
-
Downregulation of the cellular adhesion molecule Thy-1 (CD90) by cytomegalovirus infection of human fibroblasts
More LessThe deregulation of cellular adhesion molecules by human cytomegalovirus (HCMV) appears to be correlated with the development of vascular disease. In this study, it was investigated whether the expression of Thy-1 (CD90), a member of the immunoglobulin superfamily of adhesion molecules with constitutive expression on fibroblast cells, is modulated following infection with HCMV. It was observed that Thy-1 cell surface expression decreased significantly during the course of infection. Addition of neutralizing antibodies, as well as UV inactivation of virus, prevented Thy-1 downregulation. In contrast, inhibition of virus replication by cidofovir did not alter Thy-1 regulation by HCMV, indicating that immediate-early (IE) and/or early (E) gene products are responsible. Interestingly, after infection of fibroblasts with a recombinant GFP-expressing virus, infected as well as non-infected cells showed a reduced Thy-1 cell surface expression. From these findings, it is concluded that IE or E gene products of HCMV induce a so far unidentified soluble factor that mediates Thy-1 downregulation.
-
-
-
The rat cytomegalovirus homologue of parvoviral rep genes, r127, encodes a nuclear protein with single- and double-stranded DNA-binding activity that is dispensable for virus replication
More LessAn intriguing feature of the rat cytomegalovirus (RCMV) genome is open reading frame (ORF) r127, which shows similarity to the rep genes of parvoviruses as well as the U94 genes of human herpesvirus type 6A (HHV-6A) and 6B (HHV-6B). Counterparts of these genes have not been found in other herpesviruses. Here, it is shown that the r127 gene is transcribed during the early and late phases of virus replication in vitro as an unspliced 1·1 kb transcript containing the complete r127 ORF. Transcripts of r127 were also detected in various organs of RCMV-infected rats at 1 week post-infection (p.i.), but only in the salivary gland at 4 months p.i. Using rabbit polyclonal antibodies raised against the r127-encoded protein (pr127), pr127 was found to be expressed as early as 12 h p.i. within the nuclei of RCMV-infected cells in vitro. Expression of pr127 was also observed within the nuclei of cells in various organs of RCMV-infected rats at 3 weeks p.i. Moreover, pr127 was demonstrated to bind single- as well as double-stranded DNA. Finally, an RCMV r127 deletion mutant (RCMVΔr127) was generated, in which the r127 ORF was disrupted. This deletion mutant, however, was shown to replicate with a similar efficiency as wild-type RCMV (wt RCMV), both in vitro and in vivo. Taken together, it is concluded that the RCMV r127 gene encodes a nuclear protein with single- and double-stranded DNA-binding activity that is dispensable for virus replication, not only in vitro, but also during the acute phase of infection in vivo.
-
-
-
Belo Horizonte virus: a vaccinia-like virus lacking the A-type inclusion body gene isolated from infected mice
More LessHere is described the isolation of a naturally occurring A-type inclusion body (ATI)-negative vaccinia-like virus, Belo Horizonte virus (VBH), obtained from a mousepox-like outbreak in Brazil. The isolated virus was identified and characterized as an orthopoxvirus by conventional methods. Molecular characterization of the virus was done by DNA cross-hybridization using Vaccinia virus (VACV) DNA. In addition, conserved orthopoxvirus genes such as vaccinia growth factor, thymidine kinase and haemagglutinin were amplified by PCR and sequenced. All sequences presented high similarity to VACV genes. Based on the sequences, phenograms were constructed for comparison with other poxviruses; VBH clustered consistently with VACV strains. Attempts to amplify the ATI gene (ati) by PCR, currently used to identify orthopoxviruses, were unsuccessful. Results presented here suggest that most of the ati gene is deleted in the VBH genome.
-
-
-
Amino acid changes in functional domains of latent membrane protein 1 of Epstein–Barr virus in nasopharyngeal carcinoma of southern China and Taiwan: prevalence of an HLA A2-restricted ‘epitope-loss variant’
More LessFull-length sequences of the Epstein–Barr virus (EBV) gene for latent membrane protein (LMP)-1 from 22 nasopharyngeal carcinoma (NPC) biopsy specimens and 18 non-neoplastic counterparts (NPI) were determined. Relative to the B95-8 strain, the amino acid sequences of the toxic-signal and transformation domains were changed variably in NPC and NPI specimens; in contrast, no change was observed in the NF-κB (nuclear factor κB) activation domain. HLA typing revealed that 47 % of NPC and 31 % of NPI specimens were HLA A2-positive. A major A2-restricted epitope within LMP-1 (residues 125–133) was analysed. At residue 126, a change of L→F was detected in 91 % (20/22) of NPC and 67 % (12/18) of NPI specimens. In addition, a deletion at residue 126 was detected in one NPC sample from Taiwan. At residue 129, a change of M→I was observed in all samples, regardless of whether they were NPC or NPI. The changes in this peptide between NPC and NPI specimens, including mutation and deletion, are statistically significant (P<0·05). A recent report indicated that this variant sequence is recognized poorly by epitope-specific T cells. Genotyping results indicated that 96 % of NPC and 67 % of NPI samples carried a type A virus. By scanning the entire sequence of LMP-1, eight distinct patterns were identified. Detailed examination of these patterns revealed that type A strains are more prevalent in NPC than in NPI specimens and are marked by the loss of an XhoI site, the presence of a 30 bp deletion and the presence of a mutated, A2-restricted, T cell target epitope sequence. These results suggest that an EBV strain carrying an HLA A2-restricted ‘epitope-loss variant’ of LMP-1 is prevalent in NPC in southern China and Taiwan.
-
-
-
Identification of the thymidylate synthase within the genome of white spot syndrome virus
More LessThymidylate synthase (TS) (EC 2.1.1.45) is essential for the de novo synthesis of dTMP in prokaryotic and eukaryotic organisms. Within the white spot syndrome virus (WSSV) genome, an open reading frame (WSV067) that encodes a 289 amino acid polypeptide showed significant homology to all known TSs from species including mammals, plants, fungi, protozoa, bacteria and DNA viruses. In this study, WSV067 was expressed in Escherichia coli, and the purified recombinant protein showed TS activity in dUMP−folate-binding assays using ultraviolet difference spectroscopy. RT-PCR and Western blot analyses showed that WSV067 was a genuine and early gene. Phylogenetic analysis revealed that WSSV-TS was more closely related to the TSs of eukaryotes than to those from prokaryotes.
-
-
-
Structure of pre-S2 N- and O-linked glycans in surface proteins from different genotypes of hepatitis B virus
More LessThe middle-sized (M) surface proteins of hepatitis B virus (HBV) and other orthohepadnaviruses contain a conserved N-glycan in their pre-S2 domain, which is essential for the secretion of viral particles. Recently, we also found O-glycans in the pre-S2 domain of M protein from woodchuck hepatitis virus (WHV) and HBV genotype D. Since the O-glycosylation motif is not conserved in all genotypes of HBV, the glycosylation patterns of HBV genotypes A and C were analysed. Pre-S2 (glyco)peptides were released from HBV-carrier-derived HBV subviral particles by tryptic digestion, purified by reversed-phase HPLC and identified by amino acid and amino-terminal sequence analysis as well as matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS). Pre-S2 N-glycans were characterized by anion-exchange chromatography, methylation analysis and on-target sequential exoglycosidase digestions in combination with MALDI-TOF-MS, demonstrating the presence of partially sialylated diantennary complex-type oligosaccharides in all genotypes examined. Pre-S2 O-glycans were characterized by on-target sequential exoglycosidase digestions in combination with MALDI-TOF-MS. The pre-S2 domain of M protein and, to a minor extent, of L (large) protein from HBV genotype C and D was partially O-glycosylated by Neu5Ac(α2–3)Gal(β1–3)GalNAcα- or Gal(β1–3)GalNAcα-units at Thr-37 within a conserved sequence context. Genotype A, containing no Thr at position 37 or 38, was not O-glycosylated. Analytical data further revealed that M protein is mostly amino-terminally acetylated in all examined genotypes and that the terminal methionine is partially oxidized. The findings may be relevant for the secretion and the immunogenicity of HBV.
-
-
-
CD8+ T cells control corneal disease following ocular infection with herpes simplex virus type 1
More LessThe role that T cell subsets play in herpetic stromal keratitis (HSK) has been the subject of intense research efforts. While most studies implicate CD4+ T cells as the principal cell type mediating primary corneal disease, recent reports using knockout mice have suggested that both CD4+ and CD8+ T cell subsets may play integral roles in modulating the disease. Furthermore, recent studies suggest that CD8+ T cells are directly involved in maintaining virus latency in infected trigeminal ganglia. This work has addressed these discrepancies by infecting the corneas of mice lacking CD4+ and CD8+ T cells with herpes simplex virus type 1 (HSV-1) and monitoring both corneal disease and latent infection of trigeminal ganglia. Results indicated that mice lacking CD8+ T cells had more severe corneal disease than either BALB/c or B6 parental strains. In contrast, mice lacking CD4+ T cells had a milder disease than parental strains. When mice were evaluated for persistence of infectious virus, only transient differences were observed in periocular tissue and corneas. No significant differences were found in persistence of virus in trigeminal ganglia or virus reactivation from explanted ganglia. These data support the following conclusions. CD4+ T cells are not required for resistance to infection with HSV-1 and probably mediate HSK. Mice lacking CD8+ T cells do not display differences in viral loads or reactivation and thus CD8+ T cells are not absolutely required to maintain latency. Finally, CD8+ T cells probably play a protective role by regulating the immunopathological response that mediates HSK.
-
- Plant Viruses
-
-
-
Analysis of the RNA of Potato yellow vein virus: evidence for a tripartite genome and conserved 3′-terminal structures among members of the genus Crinivirus
More LessDouble-stranded RNA preparations produced from potato plants graft-inoculated with a Peruvian isolate of Potato yellow vein virus (PYVV; genus Crinivirus, family Closteroviridae) contain five RNA species denoted RNA 1, RNA 2, RNA 3, x and y of approximately 8, 5·3, 3·8, 2·0 and 1·8 kbp, respectively. The complete nucleotide sequences of PYVV RNAs 1, 2 and 3 and Northern hybridization analysis showed that PYVV RNA 1 contained the replication module and an additional open reading frame (p7), while two distinct species, RNAs 2 and 3, contain the Closteroviridae hallmark gene array. Pairwise comparisons and phylogeny of genome-encoded proteins showed that PYVV shares significant homology with other criniviruses but is most closely related to the Trialeurodes vaporariorum-vectored Cucumber yellows virus. Secondary structure prediction of the 3′-untranslated regions of all three PYVV RNAs revealed four conserved stem–loop structures and a 3′-terminal pseudoknot structure, also predicted for all fully characterized members of the genus Crinivirus and some members of the genera Closterovirus and Ampelovirus.
-
-
-
-
The coat protein of tobamovirus acts as elicitor of both L 2 and L 4 gene-mediated resistance in Capsicum
More LessIn Capsicum, the resistance conferred by the L 2 gene is effective against all of the pepper-infecting tobamoviruses except Pepper mild mottle virus (PMMoV), whereas that conferred by the L 4 gene is effective against them all. These resistances are expressed by a hypersensitive response, manifested through the formation of necrotic local lesions (NLLs) at the primary site of infection. The Capsicum L 2 gene confers resistance to Paprika mild mottle virus (PaMMV), while the L 4 gene is effective against both PaMMV and PMMoV. The PaMMV and PMMoV coat proteins (CPs) were expressed in Capsicum frutescens (L 2 L 2) and Capsicum chacoense (L 4 L 4) plants using the heterologous Potato virus X (PVX)-based expression system. In C. frutescens (L 2 L 2) plants, the chimeric PVX virus containing the PaMMV CP was localized in the inoculated leaves and produced NLLs, whereas the chimeric PVX containing the PMMoV CP infected the plants systemically. Thus, the data indicated that the PaMMV CP is the only tobamovirus factor required for the induction of the host response mediated by the Capsicum L 2 resistance gene. In C. chacoense (L 4 L 4) plants, both chimeric viruses were localized to the inoculated leaves and produced NLLs, indicating that either PaMMV or PMMoV CPs are required to elicit the L 4 gene-mediated host response. In addition, transient expression of PaMMV CP into C. frutescens (L 2 L 2) leaves and PMMoV CP into C. chacoense (L 4 L 4) leaves by biolistic co-bombardment with a β-glucuronidase reporter gene led to the induction of cell death and the expression of host defence genes in both hosts. Thus, the tobamovirus CP is the elicitor of the Capsicum L 2 and L 4 gene-mediated hypersensitive response.
-
-
-
An important determinant of the ability of Turnip mosaic virus to infect Brassica spp. and/or Raphanus sativus is in its P3 protein
More LessTurnip mosaic virus (TuMV, genus Potyvirus, family Potyviridae) infects mainly cruciferous plants. Isolates Tu-3 and Tu-2R1 of TuMV exhibit different infection phenotypes in cabbage (Brassica oleracea L.) and Japanese radish (Raphanus sativus L.). Infectious full-length cDNA clones, pTuC and pTuR1, were constructed from isolates Tu-3 and Tu-2R1, respectively. Progeny virus derived from infections with pTuC induced systemic chlorotic and ringspot symptoms in infected cabbage, but no systemic infection in radish. Virus derived from plants infected with pTuR1 induced a mild chlorotic mottle in cabbage and infected radish systemically to induce mosaic symptoms. By exchanging genome fragments between the two virus isolates, the P3-coding region was shown to be responsible for systemic infection by TuMV and the symptoms it induces in cabbage and radish. Moreover, exchanges of smaller parts of the P3 region resulted in recombinants that induced complex infection phenotypes, especially the combination of pTuC-derived N-terminal sequence and pTuR1-derived C-terminal sequence. Analysis by tissue immunoblotting of the inoculated leaves showed that the distributions of P3-chimeric viruses differed from those of the parents, and that the origin of the P3 components affected not only virus accumulation, but also long-distance movement. These results suggest that the P3 protein is an important factor in the infection cycle of TuMV and in determining the host range of this and perhaps other potyviruses.
-
-
-
Complete nucleotide sequence and genome organization of Grapevine leafroll-associated virus 3, type member of the genus Ampelovirus
More LessThis study reports on the complete genome sequence of Grapevine leafroll-associated virus 3, the type member of the genus Ampelovirus. The genome is 17 919 nt in size and contains 13 open reading frames (ORFs). Previously, the sequence of 13 154 nt of the 3′-terminal of the genome was reported. The newly sequenced portion contains a 158 nt 5′ UTR, a single papain-like protease and a methyltransferase-like (MT) domain. ORF1a encodes a large polypeptide with a molecular mass of 245 kDa. With a predicted +1 frameshift, the large fusion protein generated from ORF1a/1b would produce a 306 kDa polypeptide. Phylogenetic analysis using MT domains further supports the creation of the genus Ampelovirus for mealy-bug-transmitted viruses in the family Closteroviridae.
-
- Fungal Viruses
-
-
-
Molecular characterization of Penicillium chrysogenum virus: reconsideration of the taxonomy of the genus Chrysovirus
More LessMolecular cloning and complete nucleotide sequencing of Penicillium chrysogenum virus (PcV) dsRNAs indicated that PcV virions contained four dsRNA segments with sizes of 3562, 3200, 2976 and 2902 bp. Each dsRNA segment had unique sequences and contained a single large open reading frame (ORF). In vitro translation of transcripts derived from full-length cDNA clones of PcV dsRNAs yielded single products of sizes similar to those predicted from the deduced amino acid sequences of the individual ORFs. Sequence similarity searches revealed that dsRNA1 encodes a putative RNA-dependent RNA polymerase. In this study, it was determined that dsRNA2 encodes the major capsid protein and that p4, encoded by dsRNA4, is virion-associated as a minor component. All four dsRNAs of PcV, like the genomic segments of viruses with multipartite genomes, were found to have extended regions of highly conserved terminal sequences at both ends. In addition to the strictly conserved 5′-terminal 10 nt, a second region consisting of reiteration of the sequence CAA was found immediately upstream of the AUG initiator codon. These (CAA) n repeats are reminiscent of the translational enhancer elements of tobamoviruses. The 3′-terminal 14 nt were also strictly conserved. As PcV and related viruses with four dsRNA segments (genus Chrysovirus) have not been previously characterized at the molecular level, they were provisionally classified in the family Partitiviridae, comprising viruses with bipartite genomes. This study represents the first report on molecular characterization of a chrysovirus and the results suggest the creation of a new family of mycoviruses with multipartite dsRNA genomes to accommodate PcV and related viruses.
-
-
- Other Agents
-
-
-
Prion protein gene polymorphisms in healthy and scrapie-affected Spanish sheep
More LessThe Rasa Aragonesa sheep is the second most important Spanish breed after the Merino breed. Reported here is the prion protein (PrP) haplotype frequency distribution for scrapie-related codons (136, 154 and 171) and a sequencing study of the complete PrP gene open reading frame for this breed and six other closely related breeds. The study includes four scrapie-affected sheep flocks belonging to Rasa Aragonesa and Rasa Navarra breeds. Thirty-eight scrapie-affected sheep, 502 healthy sheep from scrapie-affected flocks and 905 sheep from a breed survey were genotyped. The most frequent PrP haplotype in both scrapie and healthy flocks was ARQ, which was found at significantly higher frequency in scrapie-affected sheep. The susceptibility-associated VRQ haplotype was found at low frequencies in six out of eight breeds, but was not present in the 38 scrapie-affected sheep. The resistance-associated ARR haplotype was found in all breeds except one (Ojinegra) at frequencies ⩾14 %. Fourteen amino acid polymorphisms were detected in these Spanish sheep, including the known amino acid substitutions at codons 112, 136, 141, 143, 154, 171 and 176, and new polymorphisms at codons 101 (Q→R), 151 (R→G), 151 (R→H), 172 (Y→D) and 175 (Q→E). Most of the novel polymorphic codons show frequencies lower than 5 %.
-
-
Volumes and issues
-
Volume 105 (2024)
-
Volume 104 (2023)
-
Volume 103 (2022)
-
Volume 102 (2021)
-
Volume 101 (2020)
-
Volume 100 (2019)
-
Volume 99 (2018)
-
Volume 98 (2017)
-
Volume 97 (2016)
-
Volume 96 (2015)
-
Volume 95 (2014)
-
Volume 94 (2013)
-
Volume 93 (2012)
-
Volume 92 (2011)
-
Volume 91 (2010)
-
Volume 90 (2009)
-
Volume 89 (2008)
-
Volume 88 (2007)
-
Volume 87 (2006)
-
Volume 86 (2005)
-
Volume 85 (2004)
-
Volume 84 (2003)
-
Volume 83 (2002)
-
Volume 82 (2001)
-
Volume 81 (2000)
-
Volume 80 (1999)
-
Volume 79 (1998)
-
Volume 78 (1997)
-
Volume 77 (1996)
-
Volume 76 (1995)
-
Volume 75 (1994)
-
Volume 74 (1993)
-
Volume 73 (1992)
-
Volume 72 (1991)
-
Volume 71 (1990)
-
Volume 70 (1989)
-
Volume 69 (1988)
-
Volume 68 (1987)
-
Volume 67 (1986)
-
Volume 66 (1985)
-
Volume 65 (1984)
-
Volume 64 (1983)
-
Volume 63 (1982)
-
Volume 62 (1982)
-
Volume 61 (1982)
-
Volume 60 (1982)
-
Volume 59 (1982)
-
Volume 58 (1982)
-
Volume 57 (1981)
-
Volume 56 (1981)
-
Volume 55 (1981)
-
Volume 54 (1981)
-
Volume 53 (1981)
-
Volume 52 (1981)
-
Volume 51 (1980)
-
Volume 50 (1980)
-
Volume 49 (1980)
-
Volume 48 (1980)
-
Volume 47 (1980)
-
Volume 46 (1980)
-
Volume 45 (1979)
-
Volume 44 (1979)
-
Volume 43 (1979)
-
Volume 42 (1979)
-
Volume 41 (1978)
-
Volume 40 (1978)
-
Volume 39 (1978)
-
Volume 38 (1978)
-
Volume 37 (1977)
-
Volume 36 (1977)
-
Volume 35 (1977)
-
Volume 34 (1977)
-
Volume 33 (1976)
-
Volume 32 (1976)
-
Volume 31 (1976)
-
Volume 30 (1976)
-
Volume 29 (1975)
-
Volume 28 (1975)
-
Volume 27 (1975)
-
Volume 26 (1975)
-
Volume 25 (1974)
-
Volume 24 (1974)
-
Volume 23 (1974)
-
Volume 22 (1974)
-
Volume 21 (1973)
-
Volume 20 (1973)
-
Volume 19 (1973)
-
Volume 18 (1973)
-
Volume 17 (1972)
-
Volume 16 (1972)
-
Volume 15 (1972)
-
Volume 14 (1972)
-
Volume 13 (1971)
-
Volume 12 (1971)
-
Volume 11 (1971)
-
Volume 10 (1971)
-
Volume 9 (1970)
-
Volume 8 (1970)
-
Volume 7 (1970)
-
Volume 6 (1970)
-
Volume 5 (1969)
-
Volume 4 (1969)
-
Volume 3 (1968)
-
Volume 2 (1968)
-
Volume 1 (1967)
Most Read This Month


