 -
Volume 88,
Issue 10,
2007
-
Volume 88,
Issue 10,
2007
Volume 88, Issue 10, 2007
- Animal
-
- RNA viruses
-
-
Immunization of Syrian golden hamsters with F subunit vaccine of human metapneumovirus induces protection against challenge with homologous or heterologous strains
More LessHuman metapneumovirus (hMPV), a newly discovered paramyxovirus, is associated with acute respiratory-tract illness, primarily in young children, individuals with underlying disease and the elderly. Two genetic lineages of hMPV circulate around the world, and viruses from these two lineages demonstrate antigenic differences. The clinical impact of hMPV warrants the development of vaccines. Recombinant soluble fusion (F) proteins of prototype viruses of the two main lineages of hMPV that can be produced in high yields have been constructed. In this study, the antigenicity, immunogenicity and protective efficacy of these soluble F subunit vaccines were evaluated in Syrian golden hamsters (Mesocricetus auratus). Immunization of hamsters with the soluble F proteins, adjuvanted with Specol or iscom matrix, induced high virus-neutralization titres, with higher titres against the homologous than the heterologous virus. The neutralizing antibodies protected from subsequent infection of the lungs with both homologous and heterologous virus. Upon challenge, viral titres in the nasal turbinates of immunized animals were reduced significantly compared with those of PBS-immunized animals. In conclusion, a soluble F subunit vaccine for hMPV that induces cross-protective immunity for infection of the lower respiratory tract in Syrian golden hamsters has been generated.
-
-
-
Measles virus minigenomes encoding two autofluorescent proteins reveal cell-to-cell variation in reporter expression dependent on viral sequences between the transcription units
More LessTranscription from morbillivirus genomes commences at a single promoter in the 3′ non-coding terminus, with the six genes being transcribed sequentially. The 3′ and 5′ untranslated regions (UTRs) of the genes (mRNA sense), together with the intergenic trinucleotide spacer, comprise the non-coding sequences (NCS) of the virus and contain the conserved gene end and gene start signals, respectively. Bicistronic minigenomes containing transcription units (TUs) encoding autofluorescent reporter proteins separated by measles virus (MV) NCS were used to give a direct estimation of gene expression in single, living cells by assessing the relative amounts of each fluorescent protein in each cell. Initially, five minigenomes containing each of the MV NCS were generated. Assays were developed to determine the amount of each fluorescent protein in cells at both cell population and single-cell levels. This revealed significant variations in gene expression between cells expressing the same NCS-containing minigenome. The minigenome containing the M/F NCS produced significantly lower amounts of fluorescent protein from the second TU (TU2), compared with the other minigenomes. A minigenome with a truncated F 5′ UTR had increased expression from TU2. This UTR is 524 nt longer than the other MV 5′ UTRs. Insertions into the 5′ UTR of the enhanced green fluorescent protein gene in the minigenome containing the N/P NCS showed that specific sequences, rather than just the additional length of F 5′ UTR, govern this decreased expression from TU2.
-
-
-
Characterization of the epitope for anti-human respiratory syncytial virus F protein monoclonal antibody 101F using synthetic peptides and genetic approaches
More LessChimeric 101F (ch101F) is a mouse–human chimeric anti-human respiratory syncytial virus (HRSV) neutralizing antibody that recognizes residues within antigenic site IV, V, VI of the fusion (F) glycoprotein. The binding of ch101F to a series of peptides overlapping aa 422–438 spanning antigenic site IV, V, VI was analysed. Residues 423–436 comprise the minimal peptide sequence for ch101F binding. Substitution analysis revealed that R429 and K433 are critical for ch101F binding, whilst K427 makes a minor contribution. Binding of ch101F to a series of single mutations at positions 427, 429 and 433 in the F protein expressed recombinantly on the cell surface confirmed the peptide results. Sequence analysis of viruses selected for resistance to neutralization by ch101F indicated that a single change (K433T) in the F protein allowed ch101F escape. The results confirm that ch101F and palivizumab have different epitope specificity and define key residues for ch101F recognition.
-
-
-
Molecular mechanisms of reversion to the ts+ (non-temperature-sensitive) phenotype of influenza A cold-adapted (ca) virus strains
More LessA ts+ ca− (non-temperature-sensitive, non-cold-adapted) revertant of the A/Leningrad/134/47/57 ca strain influenza virus [A/Leningrad/134/47/ts+18/1957(H2N2)], obtained in our previous study, lost phenotypic manifestation of ts mutations by the PB2, NP and NS genes, although, according to sequencing data, it acquired only two true reversions of a mutation in the PB2 and PB1 genes. Direct sequencing showed the appearance of 27 additional mutations (13 coding) in the genes encoding the PB2, PB1, PA, NP, M and NS proteins of the revertant, along with the above-mentioned two true reversions. We conjecture that some of these mutations suppressed phenotypic manifestation of ts mutations in the NS and NP genes.
-
-
-
Superinfection exclusion in BHK-21 cells persistently infected with Junín virus
More LessWe characterized a persistently Junín virus (JUNV)-infected BHK-21 cell line obtained by experimental infection with the XJCl3 strain. This cell line, named K3, produced low levels of virus in supernatants which were not influenced by the presence of defective interfering (DI) particles after the first year of infection. K3 cells were able to exclude superinfection of the homologous JUNV and the antigenically related Tacaribe virus (TCRV), whereas the non-related arenaviruses lymphocytic choriomeningitis virus (LCMV) and Pichinde virus (PICV) could replicate normally. Although superinfecting virus binding and internalization to persistently infected cells were slightly reduced, earlier biosynthesis of antigenomic RNA was observed in comparison with BHK-21 cells. Despite the fact that superinfection did not increase the number of cells expressing viral antigens, de novo synthesis of superinfecting virus proteins was detected. The virus produced by JUNV-superinfected K3 cells remained mostly cell-associated in the form of particles tethered to the plasma membrane and aberrant tubular structures. JUNV restriction was correlated with an overexpression of cellular protein TSG101 in K3 cells, which has been pointed out as involved in the budding of several RNA viruses. This correlation was also observed in a cell clone isolated from K3. Reduction of TSG101 expression favoured the release of infectious virus to the supernatant of JUNV-superinfected K3 cells. Our data suggest that overexpression of TSG101 in K3 cells is a novel mechanism that may contribute, along with a diminished synthesis of superinfecting virus proteins, to explain superinfection exclusion in persistently arenavirus-infected cells.
-
-
-
Persistent memory CD4+ and CD8+ T-cell responses in recovered severe acute respiratory syndrome (SARS) patients to SARS coronavirus M antigen
More LessThe membrane (M) protein of severe acute respiratory syndrome coronavirus (SARS-CoV) is a major glycoprotein with multiple biological functions. In this study, we found that memory T cells against M protein were persistent in recovered SARS patients by detecting gamma interferon (IFN-γ) production using ELISA and ELISpot assays. Flow cytometric analysis showed that both CD4+ and CD8+ T cells were involved in cellular responses to SARS-CoV M antigen. Furthermore, memory CD8+ T cells displayed an effector memory cell phenotype expressing CD45RO− CCR7− CD62L−. In contrast, the majority of IFN-γ + CD4+ T cells were central memory cells with the expression of CD45RO+ CCR7+ CD62L−. The epitope screening from 30 synthetic overlapping peptides that cover the entire SARS-CoV M protein identified four human T-cell immunodominant peptides, p21-44, p65-91, p117-140 and p200-220. All four immunodominant peptides could elicit cellular immunity with a predominance of CD8+ T-cell response. This data may have important implication for developing SARS vaccines.
-
-
-
Comparative analysis of innate immune responses following infection of newborn calves with bovine rotavirus and bovine coronavirus
More LessBovine rotavirus (BRV) and bovine coronavirus (BCV) are important causes of diarrhoea and death in newborn calves. Although these viruses belong to distinct viral classes, they both infect intestinal epithelial cells and induce similar clinical symptoms. Rotavirus usually causes an acute infection, but coronavirus infection can persist and reoccur in adults. Differences in viral structure and clinical outcome prompted us to postulate that innate mucosal immune responses would be markedly different following rotavirus and coronavirus infections. To address this hypothesis, gene expression following BRV and BCV infection was analysed in surgically prepared intestinal loops from 1-day-old colostrum-deprived calves. Gene expression was profiled at 18 h post-infection using bovine cDNA microarrays; the majority of differentially expressed significant genes were associated with the cell cycle and innate immune responses. A select group of these genes was validated by quantitative real-time PCR (qRT-PCR). The expression of genes associated with interferons (IFNs), cytokines and Toll-like receptors, which were not present on the microarray, was analysed further by qRT-PCR. Strong activation of TLR3, IL-6 and p65 was observed in BRV-infected host tissues, but not in tissues infected with BCV. Both viruses also downregulated IFN- and pro-inflammatory cytokine-associated pathways. In vitro studies confirmed that IFN inhibited viral replication. All of these results together suggested either that very early events of host responses at 18 h post-infection were being observed, or that both viruses have unique effective strategies to evade host immune responses.
-
-
-
Molecular epidemiological analyses of Japanese encephalitis virus isolates from swine in Japan from 2002 to 2004
More LessReiko Nerome, Shigeru Tajima, Tomohiko Takasaki, Takashi Yoshida, Akira Kotaki, Chang-Kweng Lim, Mikako Ito, Akira Sugiyama, Akinori Yamauchi, Takuya Yano, Taeko Kameyama, Ichiko Morishita, Masaru Kuwayama, Tomoko Ogawa, Keiji Sahara, Asaka Ikegaya, Masahiro Kanda, Yoshiyuki Hosoya, Kiyomasa Itokazu, Hajime Onishi, Seizou Chiya, Yasuko Yoshida, Yukiko Tabei, Kazuko Katsuki, Koji Tabata, Seiya Harada and Ichiro KuraneTo characterize Japanese encephalitis virus (JEV) strains recently prevalent in Japan, JEV surveillance was performed in pigs from 2002 to 2004. Eleven new JEV isolates were obtained and compared with previous isolates from Japan and other Asian countries. All of the isolates were classified into genotype 1 by nucleotide sequence analysis of the E gene. Two new isolates with different levels of neurovirulence and neuroinvasiveness, but with only one nucleotide difference in the E gene, Sw/Mie/34/2004 and Sw/Mie/40/2004, were isolated at the same farm on the same day. Sw/Mie/40/2004 displayed higher neurovirulence and neuroinvasiveness in mice than the other four new isolates. Another new isolate, Sw/Hiroshima/25/2002, was neutralized by antiserum to Beijing-1 at a level similar to the homologous Beijing-1 strain, whilst seven other new isolates were neutralized at 10-fold-lower titres. However, there were no amino acid differences in the E protein among these eight isolates. The present study indicated that the 11 new JEV isolates were genetically similar, but biologically and serologically heterogeneous.
-
-
-
Rubella virus-induced superinfection exclusion studied in cells with persisting replicons
More LessFor the first time, homologous superinfection exclusion was documented for rubella virus (RUB) by using Vero cells harbouring persisting RUB replicons. Infection with wild-type RUB was reduced by tenfold, whereas Sindbis virus infection was unaffected. Replication following infection with packaged replicons and transfection with replicon transcripts was also restricted in these cells, indicating that restriction occurred after penetration and entry. Translation of such ‘supertransfecting’ replicon transcripts was not impaired, but no accumulation of supertransfecting replicon RNA could be detected. We tested the hypothesis favoured in the related alphaviruses that superinfection exclusion is mediated by cleavage of the incoming non-structural precursor by the pre-existing non-structural (NS) protease, resulting in an inhibition of minus-strand RNA synthesis. However, cleavage of a precursor translated from a supertransfecting replicon transcript with an NS protease catalytic-site mutation was not detected and the event in the replication cycle at which superinfection exclusion is executed remains to be elucidated.
-
-
-
Increased human immunodeficiency virus type 1 Env expression and antibody induction using an enhanced alphavirus vector
More LessViral vectors encoding heterologous vaccine antigens are potent inducers of cellular immune responses, but they are generally less efficient at stimulating humoral immunity. To improve the induction of antibody responses by Semliki Forest virus-based vaccines, a vector encoding a translation-enhancer element and a novel internal signal sequence for increased expression and secretion of soluble antigens was designed. Approximately tenfold more human immunodeficiency virus type 1 gp120 was secreted into culture supernatants of infected cells using the enhanced vector compared with the parental vector. This translated into a significant increase in gp120-specific antibodies in immunized mice, suggesting that antigen-expression levels from the parental vector are limiting for induction of antibody responses. These data encourage the use of the enhanced vector for elicitation of immune responses against heterologous antigens during vaccination.
-
-
-
Effects of vpu start-codon mutations on human immunodeficiency virus type 1 replication in macrophages
More LessThe human immunodeficiency virus type 1 (HIV-1) vpu protein increases the release of virus particles from infected cells. Mutations that abrogate vpu function have a profound effect on HIV-1 replication in primary macrophage cultures. About 1.24 % of primary isolates in the HIV databases have vpu start-codon mutations. In addition, the envelope of the AD8 isolate was reported to compensate for the lack of vpu, whilst the YU-2 virus (cloned directly from the brain tissue of an infected individual) is macrophage-tropic, despite having a vpu start-codon mutation. These observations raise the possibility that envelopes evolve to compensate for the loss of vpu function in vivo. Chimeric vpu + and vpu − replication-competent clones were constructed that contained the envelopes of SF162, AD8 or YU-2. Macrophages were infected with these chimeras and virus release was measured over time by a reverse transcriptase ELISA. It was found that vpu-deficient chimeras carrying AD8 and YU-2 envelopes were consistently released at lower levels than their wild-type (wt) vpu counterparts, indicating that these envelopes did not compensate for the lack of vpu. Non-chimeric vpu + and vpu − AD8 and YU-2 followed similar patterns, although replication by vpu-deficient AD8 was variable, with virion release reaching 60 % of that recorded for AD8 with a wt vpu. In summary, no evidence was found that the AD8 or YU-2 envelopes can compensate for the lack of vpu for replication in macrophages.
-
-
-
A feline immunodeficiency virus vif-deletion mutant remains attenuated upon infection of newborn kittens
More LessThis report characterizes lentivirus attenuation associated with a vif mutation by inoculation of newborn kittens with a vif-deleted feline immunodeficiency virus provirus plasmid (FIV-pPPRΔvif). Virus in peripheral blood, antiviral antibody or CD4 T-cell count alterations were not detected in kittens inoculated with FIV-pPPRΔvif plasmid, with the exception of one kitten that demonstrated FIV Gag antibody production at 42 weeks after inoculation. In contrast, wild-type FIV-pPPR-infected kittens were viraemic, seropositive and exhibited a decrease in the CD4 T-cell subset in peripheral blood. Interestingly, FIV-specific T-cell proliferative responses detected at 32 and 36 weeks after infection were comparable for both FIV-pPPRΔvif- and wild-type FIV-pPPR-inoculated kittens and suggested the possibility of a discreet tissue reservoir supporting sustained FIV-pPPRΔvif expression or replication. Overall, these findings confirmed that the severe virus attenuation for both replication and pathogenicity exhibited by a vif-deleted FIV mutant is similar for both neonatal and adult hosts.
-
-
-
Impaired hyperphosphorylation of rotavirus NSP5 in cells depleted of casein kinase 1α is associated with the formation of viroplasms with altered morphology and a moderate decrease in virus replication
More LessThe rotavirus (RV) non-structural protein 5, NSP5, is encoded by the smallest of the 11 genomic segments and localizes in ‘viroplasms’, cytoplasmic inclusion bodies in which viral RNA replication and packaging take place. NSP5 is essential for the replicative cycle of the virus because, in its absence, viroplasms are not formed and viral RNA replication and transcription do not occur. NSP5 is produced early in infection and undergoes a complex hyperphosphorylation process, leading to the formation of proteins differing in electrophoretic mobility. The role of hyperphosphorylation of NSP5 in the replicative cycle of rotavirus is unknown. Previous in vitro studies have suggested that the cellular kinase CK1α is responsible for the NSP5 hyperphosphorylation process. Here it is shown, by means of specific RNA interference, that in vivo, CK1α is the enzyme that initiates phosphorylation of NSP5. Lack of NSP5 hyperphosphorylation affected neither its interaction with the virus VP1 and NSP2 proteins normally found in viroplasms, nor the production of viral proteins. In contrast, the morphology of viroplasms was altered markedly in cells in which CK1α was depleted and a moderate decrease in the production of double-stranded RNA and infectious virus was observed. These data show that CK1α is the kinase that phosphorylates NSP5 in virus-infected cells and contribute to further understanding of the role of NSP5 in RV infection.
-
-
-
Design of primers and use of RT-PCR assays for typing European bluetongue virus isolates: differentiation of field and vaccine strains
More LessBluetongue virus (BTV) is the causative agent of bluetongue, a disease of ruminant livestock that occurs almost worldwide between latitudes 3 ° S and 5 ° N. There are 24 serotypes of BTV (currently identified by serum neutralization assays). Since 1998, eight strains of six BTV serotypes (1, 2, 4, 8, 9 and 16) have invaded Europe. The most variable BTV protein is major outer-capsid component VP2, encoded by segment 2 (Seg-2) of the double-stranded RNA virus genome. VP2 represents the major target for neutralizing (and protective) antibodies that are generated in response to BTV infection, and is therefore the primary determinant of virus serotype. RT-PCR primers and assays targeting Seg-2 have been developed for rapid identification (within 24 h) of the six European BTV types. These assays are sensitive, specific and show perfect agreement with the results of conventional virus-neutralization methods. Previous studies have identified sequence variations in individual BTV genome segments that allow different isolates to be grouped on the basis of their geographical origins (topotypes). The assays described in this paper can detect any of the BTV isolates of the homologous serotype that were tested from different geographical origins (different Seg-2 topotypes). Primers were also identified that could be used to distinguish members of these different Seg-2 topotypes, as well as field and vaccine strains of most of the European BTV serotypes. The serotype-specific assays (and primers) showed no cross-amplification when they were evaluated with multiple isolates of the most closely related BTV types or with reference strains of the remaining 24 serotypes. Primers developed in this study will be updated periodically to maintain their relevance to current BTV distribution and epidemiology (http://www.iah.bbsrc.ac.uk/dsRNA_virus_proteins/ReoID/rt-pcr-primers.htm).
-
-
-
Evidence for functional significance of the permuted C motif in Co2+-stimulated RNA-dependent RNA polymerase of infectious bursal disease virus
More LessSegment B of bisegmented infectious bursal disease virus (IBDV) encodes virus protein 1 (VP1), possessing RNA-dependent RNA polymerase (RdRp) activity. This multidomain protein includes an RdRp domain with a non-canonical order of three sequence motifs forming the active site: C–A–B. The A–B–C order of the motifs, as found in RdRps of the majority of viruses, was converted by relocation (permutation) of motif C to a C–A–B order. Due to the unusual location and unproven significance, the motif was named ‘C?’. This motif includes an Ala–Asp–Asn tripeptide that replaces the C motif Gly–Asp–Asp sequence, widely considered a hallmark of RdRps. In this study, functional significance of the C? motif was investigated by using purified His-tagged VP1 mutants with either a double replacement (ADN to GDD) or two single-site mutants (ADD or GDN). All mutants showed a significant reduction of RdRp activity in vitro, in comparison to that of VP1. Only the least-affected GDN mutant gave rise to viable, albeit partially impaired, progeny using a reverse-genetics system. Experiments performed to investigate whether the C motif was implicated in the control of metal dependence revealed that, compared with Mn2+ and Mg2+, Co2+ stimulated RdRp unconventionally. No activity was observed in the presence of several divalent cations. Of two Co2+ salts with Cl− and
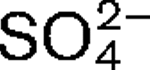 anions, the former was a stronger stimulant for RdRp. When cell-culture medium was supplemented with 50 μM Co2+, an increase in IBDV progeny yield was observed. The obtained results provide evidence that the unusual Co2+ dependence of the IBDV RdRp might be linked to the permuted organization of the motif.
anions, the former was a stronger stimulant for RdRp. When cell-culture medium was supplemented with 50 μM Co2+, an increase in IBDV progeny yield was observed. The obtained results provide evidence that the unusual Co2+ dependence of the IBDV RdRp might be linked to the permuted organization of the motif.
-
-
-
Ectropis obliqua picorna-like virus IRES-driven internal initiation of translation in cell systems derived from different origins
More LessEctropis obliqua picorna-like virus (EoPV) is an insect RNA virus that causes a lethal granulosis infection of larvae of the tea looper (Ectropis obliqua). An internal ribosome entry site (IRES) mediates translation initiation of EoPV RNA. Here, bicistronic constructs were used to examine the 5′ untranslated region (UTR) of EoPV for IRES activity. The capacities of the EoPV 5′ UTR IRES and another insect virus IRES, the cricket paralysis virus intergenic region IRES, to mediate internal translation initiation in a variety of translation systems were also compared. The results demonstrated that the EoPV IRES functioned efficiently not only in mammalian cell-derived systems, but also in an insect cell-derived translation system. However, it functioned inefficiently in a plant cell-derived translation system. This study reveals the host preferences of the EoPV IRES and important differences in IRES function between the EoPV IRES and other characterized picorna-like insect viral IRESs.
-
- DNA viruses
-
-
Comparison of the antiviral potentials among the pseudorabies-resistant transgenes encoding different soluble forms of porcine nectin-1 in transgenic mice
More LessNectin-1 is an alphaherpesvirus receptor that binds to virion glycoprotein D by the first immunoglobulin (Ig)-like domain. The possibility of making animals resistant to pseudorabies virus (PRV) infection has been investigated by generating transgenic mice expressing soluble forms of porcine nectin-1. Previously, transgenic mice were generated that expressed a fusion protein made of the entire ectodomain of nectin-1 fused to the Fc portion of human IgG, or the first Ig-like domain fused to the Fc portion of porcine IgG. Here, the contribution of the second and third Ig-like domains of nectin-1 was analysed by generating transgenic mice expressing the entire ectodomain of nectin-1 fused to the porcine Fc portion. Transgenic mice expressing each of three different fusion proteins were challenged with PRV for comparison of their resistance. Altogether, mice transgenic for a chimera that carried the entire ectodomain were more resistant than those transgenic for a chimera that carried the first Ig-like domain.
-
-
-
Cytomegaloviral proteins pUL50 and pUL53 are associated with the nuclear lamina and interact with cellular protein kinase C
More LessHuman cytomegalovirus-encoded pUL50 and pUL53 belong to a group of conserved herpesviral nuclear proteins. This study describes: (i) the co-localization of pUL50 with components of the nuclear lamina such as lamins A/C and lamin B receptor by double immunofluorescent staining, (ii) a strong pUL50-mediated relocalization of pUL53 from a diffuse nuclear pattern towards a nuclear rim localization, (iii) a direct interaction between pUL50 and pUL53, as well as between pUL50 and protein kinase C (PKC), shown by yeast two-hybrid and co-immunoprecipitation analyses, (iv) in vitro phosphorylation of pUL50, which is highly suggestive of PKC activity, and finally (v) partial relocalization of PKC by pUL50/pUL53 from its main cytoplasmic localization to a marked nuclear lamina accumulation. These data suggest a role for pUL50 and pUL53 in the recruitment of PKC, an event that is considered to be important for cytomegalovirus-induced distortion of the nuclear lamina.
-
-
-
Discovery of herpesviruses in bats
More LessSeven novel gammaherpesviruses (GHV) and one novel betaherpesvirus were discovered in seven different European bat species (order Chiroptera, family Vespertilionidae) with a pan-herpesvirus PCR assay, targeting the DNA polymerase (DPOL) gene. The sequences of six bat GHV were similarly related to members of the gammaherpesvirus genera Percavirus and Rhadinovirus. The seventh GHV was related to the porcine lymphotropic herpesvirus 1 (genus Macavirus). The betaherpesvirus appeared to be a distant relative of human cytomegalovirus. For three bat GHV a 3.6 kbp locus was amplified and sequenced, spanning part of the glycoprotein B gene and the majority of the DPOL gene. In phylogenetic analysis, the three bat GHV formed a separate clade with similar distance to the Percavirus and Rhadinovirus clades. These novel viruses are the first herpesviruses to be described in bats.
-
-
-
Post-translational modifications of Epstein–Barr virus BARF1 oncogene-encoded polypeptide
More LessEpstein–Barr virus is associated with several human lymphomas and carcinomas, and its BARF1 oncogene encodes a protein that is thought to play an important role in carcinogenesis. A BARF1 recombinant adenovirus expression system, which led us to discover the macromolecular size of the cleaved and secreted form of the BARF1 protein in the native state and its mitogenic capacity on various cell lines in culture, was used further to investigate the structure and maturation of the BARF1 protein. We recently reported biophysical studies that showed dimer-based oligomerization of the BARF1 polypeptide. Here, new data are presented that confirm post-translational modifications predicted from the BARF1 sequence: phosphorylation on serine and threonine, and N- and O-glycosylation. The N- and O-glycans were partially characterized and it was demonstrated that both modifications are required for active secretion of the BARF1 protein via the classical pathway.
-
Volumes and issues
-
Volume 105 (2024)
-
Volume 104 (2023)
-
Volume 103 (2022)
-
Volume 102 (2021)
-
Volume 101 (2020)
-
Volume 100 (2019)
-
Volume 99 (2018)
-
Volume 98 (2017)
-
Volume 97 (2016)
-
Volume 96 (2015)
-
Volume 95 (2014)
-
Volume 94 (2013)
-
Volume 93 (2012)
-
Volume 92 (2011)
-
Volume 91 (2010)
-
Volume 90 (2009)
-
Volume 89 (2008)
-
Volume 88 (2007)
-
Volume 87 (2006)
-
Volume 86 (2005)
-
Volume 85 (2004)
-
Volume 84 (2003)
-
Volume 83 (2002)
-
Volume 82 (2001)
-
Volume 81 (2000)
-
Volume 80 (1999)
-
Volume 79 (1998)
-
Volume 78 (1997)
-
Volume 77 (1996)
-
Volume 76 (1995)
-
Volume 75 (1994)
-
Volume 74 (1993)
-
Volume 73 (1992)
-
Volume 72 (1991)
-
Volume 71 (1990)
-
Volume 70 (1989)
-
Volume 69 (1988)
-
Volume 68 (1987)
-
Volume 67 (1986)
-
Volume 66 (1985)
-
Volume 65 (1984)
-
Volume 64 (1983)
-
Volume 63 (1982)
-
Volume 62 (1982)
-
Volume 61 (1982)
-
Volume 60 (1982)
-
Volume 59 (1982)
-
Volume 58 (1982)
-
Volume 57 (1981)
-
Volume 56 (1981)
-
Volume 55 (1981)
-
Volume 54 (1981)
-
Volume 53 (1981)
-
Volume 52 (1981)
-
Volume 51 (1980)
-
Volume 50 (1980)
-
Volume 49 (1980)
-
Volume 48 (1980)
-
Volume 47 (1980)
-
Volume 46 (1980)
-
Volume 45 (1979)
-
Volume 44 (1979)
-
Volume 43 (1979)
-
Volume 42 (1979)
-
Volume 41 (1978)
-
Volume 40 (1978)
-
Volume 39 (1978)
-
Volume 38 (1978)
-
Volume 37 (1977)
-
Volume 36 (1977)
-
Volume 35 (1977)
-
Volume 34 (1977)
-
Volume 33 (1976)
-
Volume 32 (1976)
-
Volume 31 (1976)
-
Volume 30 (1976)
-
Volume 29 (1975)
-
Volume 28 (1975)
-
Volume 27 (1975)
-
Volume 26 (1975)
-
Volume 25 (1974)
-
Volume 24 (1974)
-
Volume 23 (1974)
-
Volume 22 (1974)
-
Volume 21 (1973)
-
Volume 20 (1973)
-
Volume 19 (1973)
-
Volume 18 (1973)
-
Volume 17 (1972)
-
Volume 16 (1972)
-
Volume 15 (1972)
-
Volume 14 (1972)
-
Volume 13 (1971)
-
Volume 12 (1971)
-
Volume 11 (1971)
-
Volume 10 (1971)
-
Volume 9 (1970)
-
Volume 8 (1970)
-
Volume 7 (1970)
-
Volume 6 (1970)
-
Volume 5 (1969)
-
Volume 4 (1969)
-
Volume 3 (1968)
-
Volume 2 (1968)
-
Volume 1 (1967)
Most Read This Month


